Molecular orbitals¶
Visualization¶
The OrbitalViewer provides an interactive interface for visualizing molecular orbitals represented by a VeloxChem molecular orbital object.
Given a set of molecular orbitals, the viewer constructs three‑dimensional isodensity surfaces corresponding to selected orbitals and enables interactive inspection of their spatial characteristics. The OrbitalViewer operates directly on the orbital coefficients and basis information stored in the VeloxChem molecular orbital data structure, ensuring consistency with the underlying electronic‑structure calculation.
import veloxchem as vlx
molecule = vlx.Molecule.read_name("caffeine")
basis = vlx.MolecularBasis.read(molecule, "6-31g")
scf_drv = vlx.ScfRestrictedDriver()
scf_results = scf_drv.compute(molecule, basis)
orb_viewer = vlx.OrbitalViewer()
orb_viewer.visualize(molecule, basis, scf_drv.mol_orbs)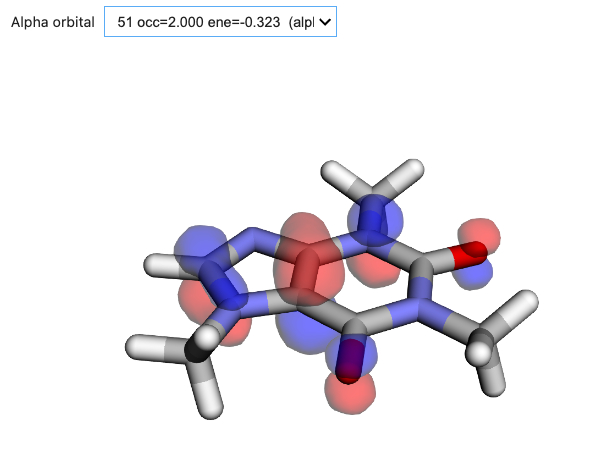
Figure: Interactive exploration of molecular orbitals is available through the OrbitalViewer. Here we show a representative isodensity surface generated with VeloxChem.
Localization¶
Orbital localization provides an alternative representation of the electronic structure in which molecular orbitals are transformed to maximize their spatial compactness or chemical interpretability, while preserving the span of the underlying orbital subspace. Localized orbitals are widely used for qualitative chemical analysis, construction of local correlation methods, and interpretation of bonding patterns in complex molecular systems.
In VeloxChem, localization is performed through unitary rotations of selected orbitals obtained from a prior electronic structure calculation. The default selection is the set of occupied orbitals, but it can alternatively be set with the keyword argument mo_range of the compute method.
The program implements the Boys and Pipek–Mezey localization schemes, which respectively optimize spatial localization and atomic charge separation criteria. Pipek–Mezey is the default choice but it can be changed to “boys” with the method attribute. For Pipek–Mezey in addition, a choice of projector can be made; the alternative to the default setting of “mulliken” is “lowdin”.
import veloxchem as vlx
molecule = vlx.Molecule.read_name("ethylene")
basis = vlx.MolecularBasis.read(molecule, "cc-pvdz")
scf_drv = vlx.ScfRestrictedDriver()
scf_results = scf_drv.compute(molecule, basis)
loc_drv = vlx.OrbitalLocalizationDriver()
loc_drv.method = "pm" # this is default
loc_drv.pm_projector = "mulliken" # this is default
loc_results = loc_drv.compute(molecule, basis, scf_results)The results dictionary of the orbital localization driver contains the localized orbitals in the form of a VeloxChem object that can be passed to the orbital viewer for visual inspection.
orb_viewer = vlx.OrbitalViewer()
orb_viewer.visualize(molecule, basis, loc_results["loc_orbs"])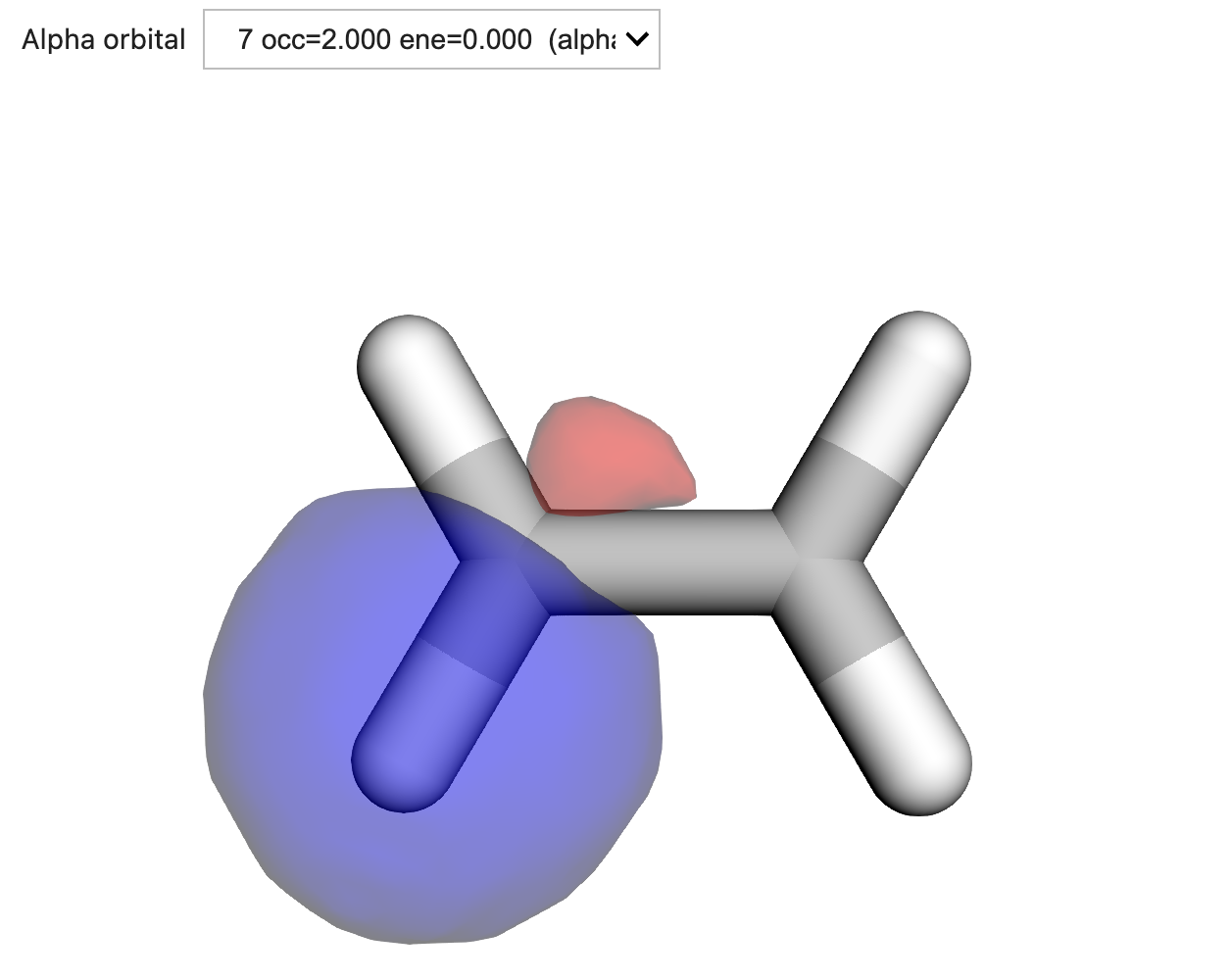
Figure: One of the occupied localized sp2-hybridized orbitals resulting from a localization performed with the Pipek–Mezey scheme using the Mulliken projector.