VeloxChem implements both implicit (CPCM and SMD) and explicit (polarizable embedding) solvation models.
Implicit solvation¶
Implicit solvation models describe the effect of a surrounding liquid environment by replacing the explicit solvent molecules with a continuous dielectric medium that interacts self‑consistently with the electronic structure of the solute. Tomasi et al. (2005)
A separation is made between equilibrium and non-equilibrium solvation. In the former case, the timescale is such that both nuclear and electronic relaxations take place in the environment, such as in molecular structure optimizations. In the latter case, only electrons are fully equilibrated with the time-dependent solute charge density, such as in UV/vis spectrum simulations.
CPCM¶
In the conductor‑like polarizable continuum model (CPCM), the solute is placed inside a cavity defined by its molecular surface, and the reaction field is obtained by solving surface‑charge equations that approximate the dielectric screening of a perfect conductor and are subsequently scaled to represent the desired solvent permittivity.
VeloxChem implements the CPCM model for:
SCF energies
gradients (structure optimizations)
linear response (UV/vis spectra and more)
Python script
import veloxchem as vlx
molecule = vlx.Molecule.read_name("ammonia")
basis = vlx.MolecularBasis.read(molecule, "def2-svp")
scf_drv = vlx.ScfRestrictedDriver()
scf_drv.xcfun = "b3lyp"
scf_drv.solvation_model = "cpcm"
scf_results = scf_drv.compute(molecule, basis)
rsp_drv = vlx.LinearResponseEigenSolver()
rsp_drv.nstates = 10
rsp_results = rsp_drv.compute(molecule, basis, scf_results)
opt_drv = vlx.OptimizationDriver(scf_drv)
opt_results = opt_drv.compute(molecule, basis, scf_results)In CPCM, the induced charge density on the cavity (due to the presence of the solute) is represented on a surface grid and discretized as Gaussian charges centred at the grid points. The converged grid charges can be visualized using the CPCM driver.
opt_molecule = opt_results["final_molecule"]
scf_drv.cpcm_drv.visualize_cpcm_charges(opt_molecule)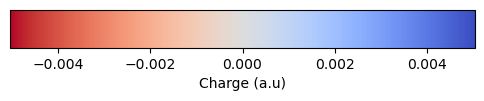
Text file
@jobs
task: scf
@end
@method settings
basis: def2-svp
xcfun: b3lyp
solvation model: cpcm
cpcm epsilon : 78.39
@end
@molecule
charge: 0
multiplicity: 1
xyz:
...
@endSMD¶
The solvation model based on density (SMD) combines a self‑consistent reaction‑field description of the electrostatic polarization with empirically parametrized terms for cavitation, dispersion, and solvent–solute interactions based on the solute’s electron density, enabling accurate free‑energy predictions across a wide range of solvents. For further details, see Marenich et al. (2009).
In Veloxchem, the electrostatic contribution is determined using the CPCM model.
Python script
import veloxchem as vlx
molecule = vlx.Molecule.read_name("methanol")
basis = vlx.MolecularBasis.read(molecule, "def2-svp")
scf_drv = vlx.ScfRestrictedDriver()
scf_drv.solvation_model = "smd"
scf_drv.smd_solvent = "water"
scf_results = scf_drv.compute(molecule, basis)Text file
@jobs
task: scf
@end
@method settings
basis: def2-svp
xcfun: b3lyp
solvation model: smd
smd solvent : water
@end
@molecule
charge: 0
multiplicity: 1
xyz:
...
@endPolarizable and electrostatic embedding¶
An explicit representation of the environment is available with:
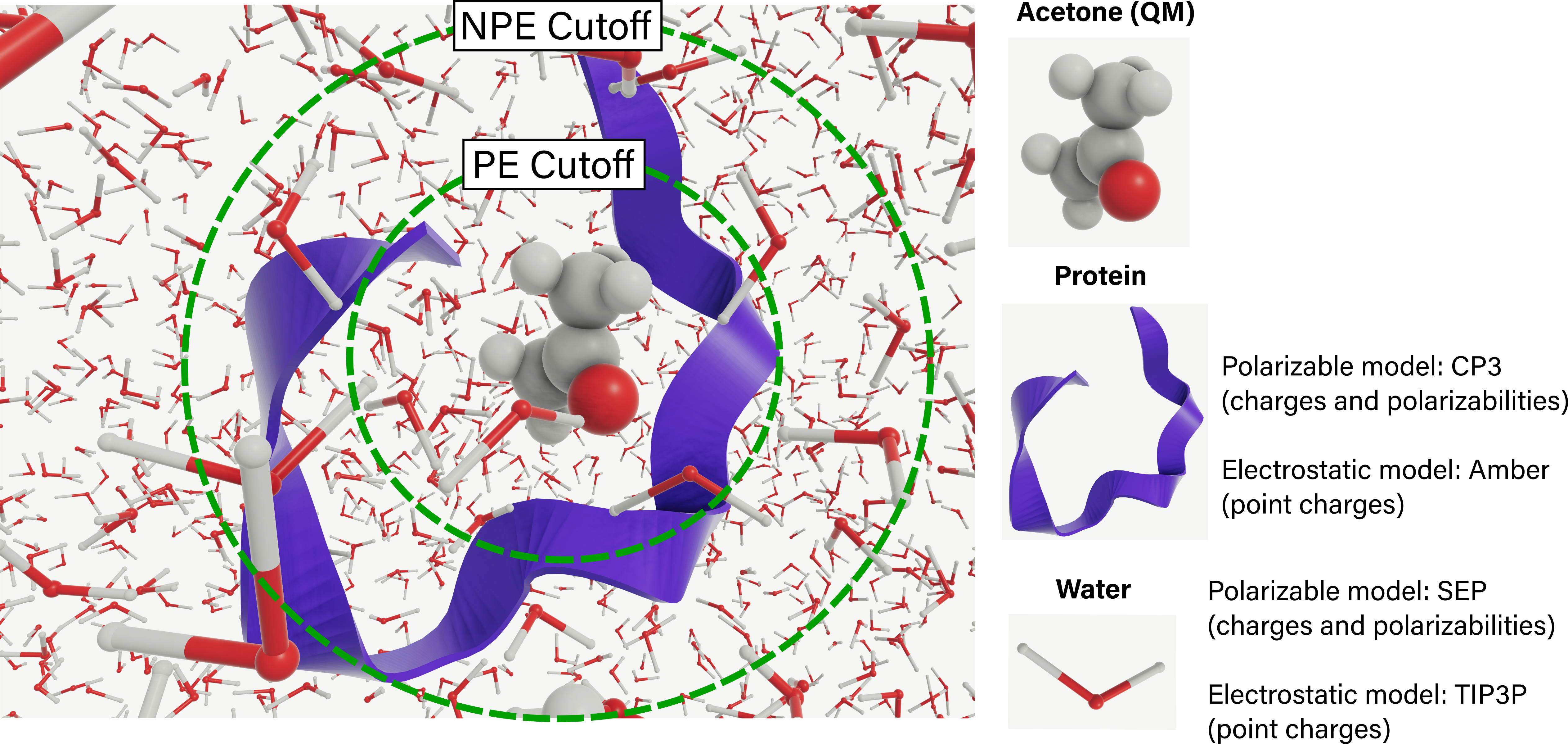
Polarizable embedding (PE), where molecules in the environment are represented by site charges and polarizabilities.
Non-polarizable (NPE), where the environment is represented by point charges only.
Python script
Either a single PDB, containing one or more structures, or a trajectory can be passed to the EnsembleParser. In this step, the PE and NPE cutoffs are selected:
import veloxchem as vlx
ens_parser = vlx.EnsembleParser()
ensemble = ens_parser.structures(
trajectory_file = "../input_files/alpha-helix-acetone-water.xtc",
topology_file = "../input_files/alpha-helix-acetone-water.tpr",
qm_region = 'resname LIG',
num_snapshots = 3,
pe_cutoff = 3.0,
npe_cutoff = 5.0,
)When providing a time-resolved trajectory, for example, .xtc together with .tpr, several options can be specified:
num_snapshots: Controls snapshot extraction.Default: All snapshots used.
If provided: Snapshots are selected evenly spaced.
start: Start time of the trajectory window in ps. Default: Time of the first snapshot.end: End time of the trajectory window in ps. Default: Time of the last snapshot.last_snapshot_only = True: Processes only the final snapshot.
Key parameters are the following:
qm_region: MDAnalysis selection string that defines QM region.qm_charge: charge of the QM regionenv_region: MDAnalysis selection string that defines environment. Ifenv_regionis not defined, the entire system is treated as QM.Default: All atoms not included in qm_region.
The number of residues treated with PE and NPE can be accessed as follows:
print("number residues PE = ", ensemble[0]["number_residues_pe"])
print("number residues NPE = ", ensemble[0]["number_residues_npe"])number residues PE = 9
number residues NPE = 14
Once the environment has been defined, the environment models can be selected with the set_env_models method of the EnsembleDriver:
ens_drv = vlx.EnsembleDriver()
ens_drv.set_env_models(
pe_model=["CP3", "SEP"],
npe_model=["ff19sb", "tip3p"],
)VeloxChem supports the following environment models for PE and NPE:
PE:
CP3Reinholdt et al. (2020) for proteins, andSEPBeerepoot et al. (2016) for common polar and non-polar solvent molecules and ions.NPE:
ff19sbTian et al. (2020) for proteins, andtip3pJorgensen et al. (1983) for water.
The SCF calculations can be automated by passing scf_options and, optionally, property_options to the compute method:
scf_options = {
"scf_type": "restricted",
"conv_thresh": 1.0e-6,
"max_iter": 150,
"xcfun": "cam-b3lyp",
"grid_level": 4,
}
property_options = {
"property": "absorption",
"nstates": 5,
"nto": True,
}results = ens_drv.compute(
ensemble,
basis_set = "aug-pcseg-1",
scf_options = scf_options,
property_options = property_options,
)In some situations, for example when preparing input files for running on a cluster, it is convenient to use the write_pot_files method to generate the potential files:
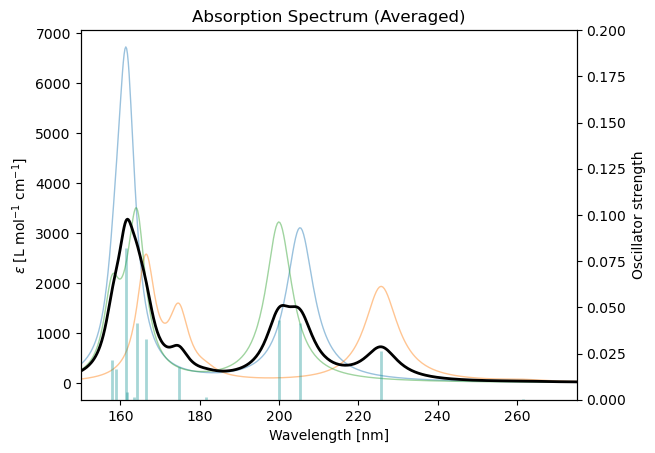
ens_drv.write_pot_files(ensemble)Finally, the averaged spectra can be plotted:
ens_drv.plot_uv_vis_spectra(
results,
show_individual = True,
show_sticks = True,
xlim_nm = (150, 275)
)
Text file
@jobs
task: scf
@end
@method settings
xcfun: cam-b3lyp
basis: aug-pcseg-1
potfile: pe_frame_000000.pot
@end
@molecule
charge: 0
multiplicity: 1
xyz:
H 23.09239578 21.92239571 22.43439484
C 23.3823967 22.95239449 22.65439606
H 24.33239555 23.14239693 22.14439583
H 22.6023941 23.562397 22.18439484
C 23.44239616 23.26239395 24.09439468
O 23.01239586 22.44239616 24.92439651
C 24.08239555 24.58239555 24.50439644
H 24.36239624 24.5623951 25.55439568
H 24.92239571 24.80239487 23.84439659
H 23.46239662 25.46239471 24.35439491
@endThe potential file named pe_frame_000000.pot in this example takes the following form: pe_frame_000000.pot
- Tomasi, J., Mennucci, B., & Cammi, R. (2005). Quantum Mechanical Continuum Solvation Models. Chem. Rev., 105(8), 2999–3094. 10.1021/cr9904009
- Marenich, A. V., Cramer, C. J., & Truhlar, D. G. (2009). Universal Solvation Model Based on Solute Electron Density and on a Continuum Model of the Solvent Defined by the Bulk Dielectric Constant and Atomic Surface Tensions. J. Phys. Chem. B, 113(18), 6378–6396. 10.1021/jp810292n
- Reinholdt, P., Kjellgren, E. R., Steinmann, C., & Olsen, J. M. H. (2020). Cost-Effective Potential for Accurate Polarizable Embedding Calculations in Protein Environments. Journal of Chemical Theory and Computation, 16(2), 1162–1174. 10.1021/acs.jctc.9b00616
- Beerepoot, M. T. P., Steindal, A. H., List, N. H., Kongsted, J., & Olsen, J. M. H. (2016). Averaged Solvent Embedding Potential Parameters for Multiscale Modeling of Molecular Properties. Journal of Chemical Theory and Computation, 12(4), 1684–1695. 10.1021/acs.jctc.5b01000
- Tian, C., Kasavajhala, K., Belfon, K. A. A., Raguette, L., Huang, H., Migues, A. N., Bickel, J., Wang, Y., Pincay, J., Wu, Q., & Simmerling, C. (2020). ff19SB: Amino-Acid-Specific Protein Backbone Parameters Trained against Quantum Mechanics Energy Surfaces in Solution. Journal of Chemical Theory and Computation, 16(1), 528–552. 10.1021/acs.jctc.9b00591
- Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W., & Klein, M. L. (1983). Comparison of simple potential functions for simulating liquid water. The Journal of Chemical Physics, 79(2), 926–935. 10.1063/1.445869