In a Jupyter notebook¶
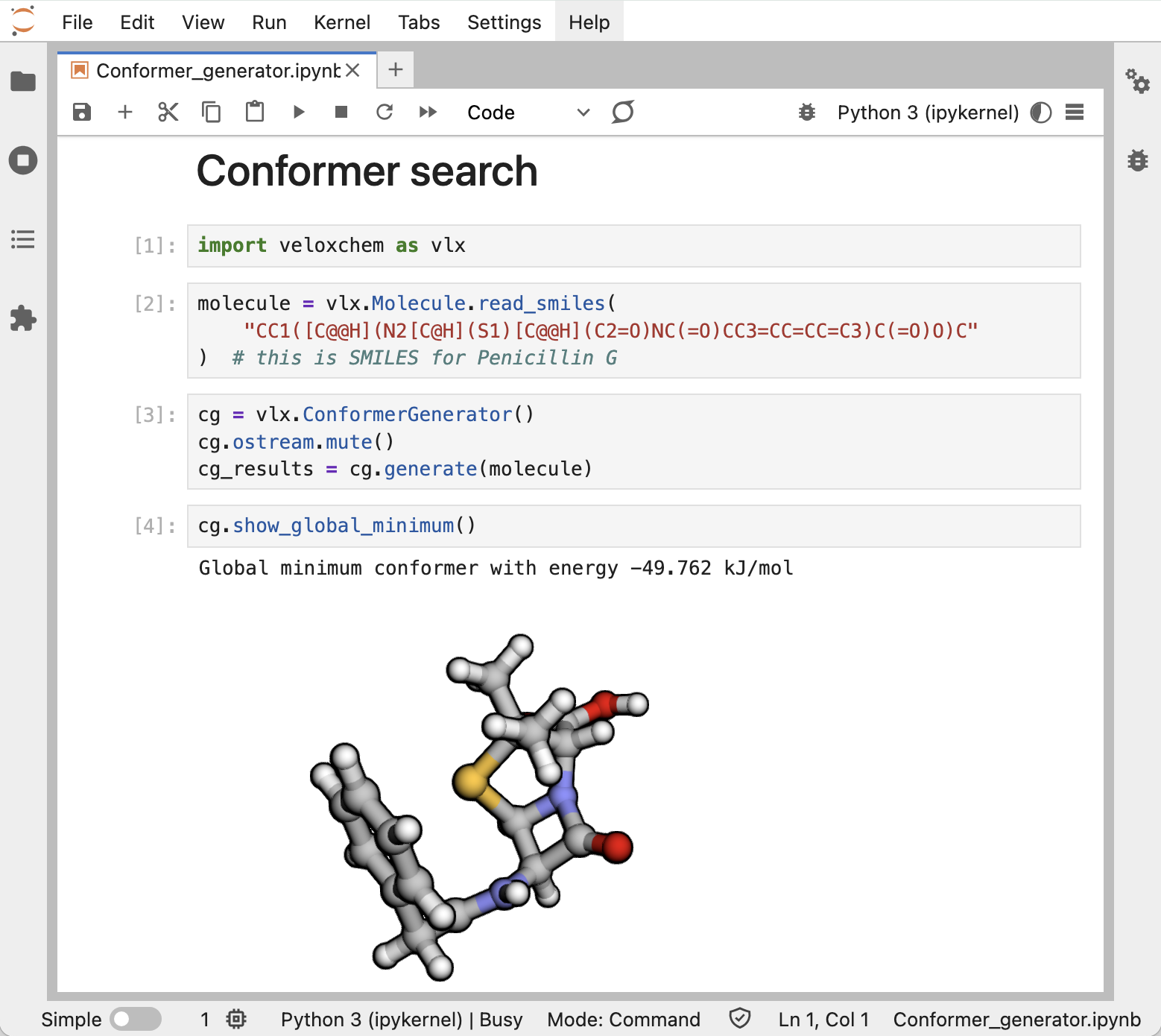
On a personal computer, we recommend importing the VeloxChem Python module and running calculations in Jupyter notebooks. This provides a very flexible framework for the creation of workflows, as amply illustrated in the eChem book Fransson et al. (2022).
Using an input file¶
Calculations can also be run in the terminal window using input files in the form of Python scripts or text files.
Python script
Terminal command:
python myjob.py > myjob.outThe Python script input file named myjob.py above can e.g. take the form:
import veloxchem as vlx
xyz_string = """
3
water
O 0.0000000 0.0000000 -0.1653507
H 0.7493682 0.0000000 0.4424329
H -0.7493682 0.0000000 0.4424329
"""
molecule = vlx.Molecule.read_xyz_string(xyz_string)
basis = vlx.MolecularBasis.read(molecule, "def2-svp")
scf_drv = vlx.ScfRestrictedDriver()
scf_drv.xcfun = "b3lyp"
scf_drv.filename = "vlx_results_hdf5"
scf_results = scf_drv.compute(molecule, basis)The results of the calculation are stored in an HDF5 file with a user specified name. This file can be directly read and analyzed with VIAMD.
Text file
Terminal command:
vlx myjob.inp [myjob.out]An input file in text format consists of multiple groups marked with @group name and @end. The default unit for Cartesian coordinates of atoms is Angstrom. An example of the input file named myjob.inp above reads:
@jobs
task: scf
@end
@method settings
xcfun: b3lyp
basis: def2-svp
@end
@molecule
charge: 0
multiplicity: 1
xyz:
O 0.00000 0.00000 0.00000
H 0.00000 0.00000 1.79524
H 1.69319 0.00000 -0.59904
@end- Fransson, T., Delcey, M., Brumboiu, I. E., Hodecker, M., Li, X., Rinkevicius, Z., Dreuw, A., Rhee, Y. M., & Norman, P. (2022). Computational Chemistry from Laptop to HPC: A notebook exploration of quantum chemistry. KTH Royal Institute of Technology. 10.30746/978-91-988114-0-7